The Full Picture
How AI is transforming medical imaging
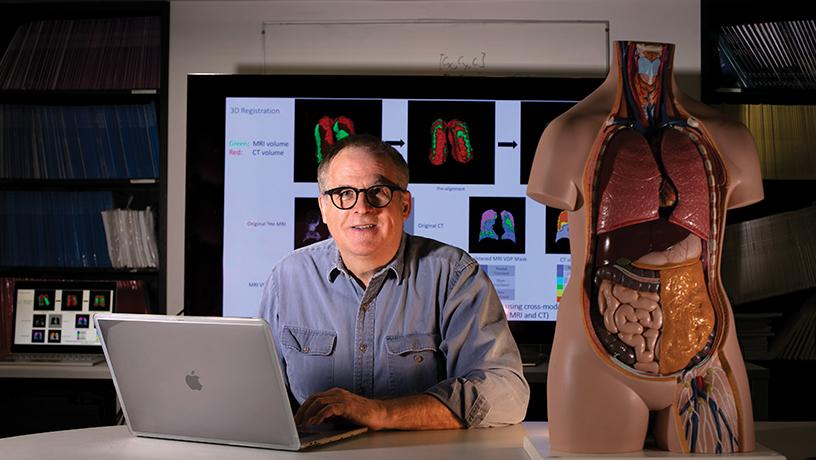
Andrew F. Laine, Percy K. and Vida L. W. Hudson Professor of Biomedical Engineering and Professor of Radiology (Physics).
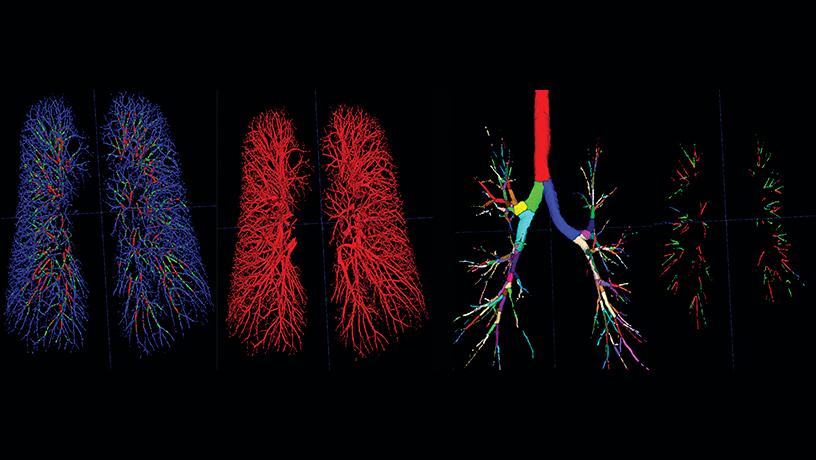
Left: 3-D rendering of pulmonary blood vessels segmented from thoracic high-resolution computed tomography.
Right: 3-D rendering of a human airway tree segmented from thoracic high-resolution computed tomography, with anatomic labeling.
Medical imaging technologies can save countless lives, but only when doctors and researchers have the right tools to mine those images for insight. Using artificial intelligence, engineers are increasingly enabling fast, convenient, accurate observations of how the body operates—and treatments when we inevitably fall ill.
Sometimes that AI can upend conventional wisdom. For instance, physicians long thought there were three types of emphysema related to chronic obstructive pulmonary disease (COPD), an inflammatory lung condition and the third-leading cause of death worldwide. It’s not well understood and there’s no known cure, “so it’s essentially a death sentence,” says Andrew F. Laine, professor of biomedical engineering and of radiology (Physics).
But when radiologists told him they had a hunch that more than three types existed, Laine embarked on a decade-long study of longitudinal databases containing lung CT images of healthy and sick people. Using deep learning to tease out textural correlations (later tied to clinical observations), his lab has identified nine new subtypes of emphysema, a distinction that could deepen understanding of the disease’s physiological stages and provide individualized pathways for treatment.
Another COPD mystery is whether it’s a vascular disease, caused by inflammation of blood vessels in the lungs, or an airway disease, in which cells of the airway passages degrade. Laine believes it’s possibly both. His team, in collaboration with Dr. R. Graham Barr (Medicine) and Elsa D. Angelini (Radiology) of Columbia University Irving Medical Center, has found three vascular genes associated with newly discovered subtypes. He’s now examining individual differences in airway tree structure associated with COPD risk. “The airway trees are almost like fingerprints,” Laine says. “Think of it as a hickory or an oak or a birch tree, each with its distinct branching structure.” Through models guided by known early biological development stages of the lung, deep learning might help explain why some people are more susceptible to diseases associated with smoking or air pollution. Eventually, knowing individual tree subtypes could lead to a better understanding of risk factors.
This is a new field called few-shot learning. When you very quickly want to generalize intelligence to a new domain, how do you teach a machine to learn with just a few new samples?

Shih-Fu Chang, Richard Dicker Professor of Electrical Engineering and professor of computer science, and Samuel K. Sia, professor of biomedical engineering.
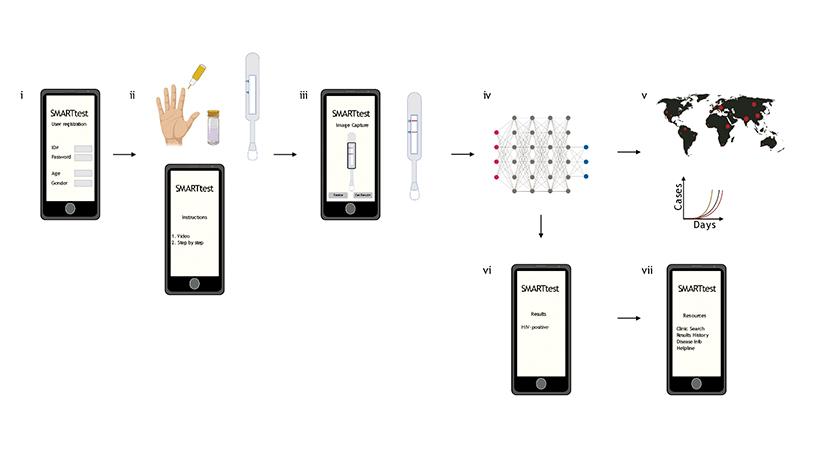
Digital platform for ensuring reliable interpretation of lateral flow assays in decentralized settings, including front-end visualization, app scanning, and tailored follow-up instructions.
Infectious lung diseases like COVID-19 create special urgency, and medical imaging has a huge role to play in bringing the pandemic under control—notably, in quickly and seamlessly diagnosing people at home. Some test kits, called lateral-flow assays, operate like pregnancy tests. Apply a fluid (in this case blood) to one end of a plastic stick, it mixes with colored nanoparticles that bind to a molecule of interest (such as COVID-19 antibodies), and the appearance of a colored band indicates the target is present. Samuel K. Sia, professor of biomedical engineering, developed a smartphone app that takes a picture of the assay and indicates whether the band is present. It sounds simple, but the band can be faint and lighting conditions vary, so the system relies on deep learning to process the image.
Historically, training deep learning algorithms has required tens of thousands of validated images; in a pandemic, that’s often unavailable in a timely fashion. So Shih-Fu Chang, professor of electrical engineering and computer science, applied an innovation in machine learning called transfer learning. He trained Sia’s new kit using a dataset from a different test kit that he could then quickly adapt to the new target kit using a mere 10 validated images. The result: They can rapidly adapt their model to new kits with 96%–100% accuracy and are awaiting FDA approval. “This is a new field called few-shot learning,” Chang says. “When you very quickly want to generalize intelligence to a new domain, how do you teach a machine to learn with just a few new samples?”
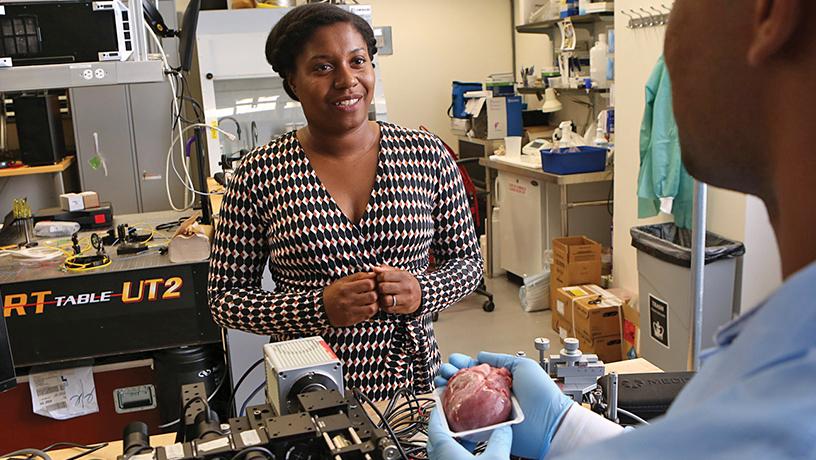
Christine P. Hendon, associate professor of electrical engineering.
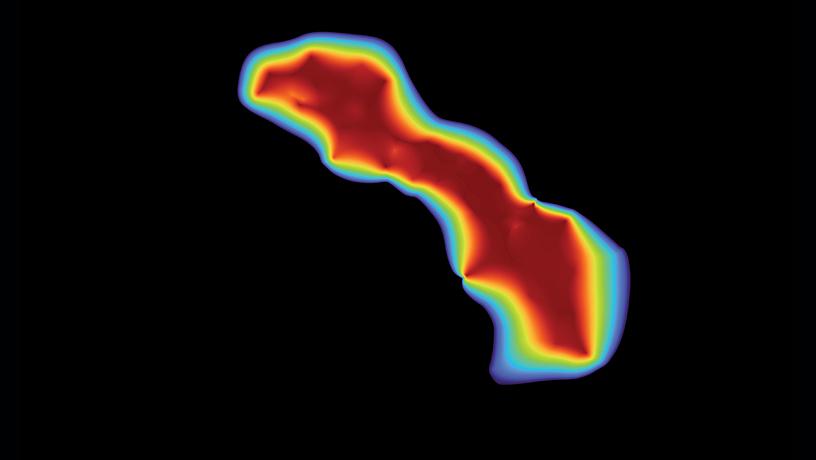
Optical reflectance spectral analysis with AI to measure ablation lesion dimensions to treat cardiac arrhythmias. Areas without lesions are shown in blue, lesions in red.
Ultimately, imaging hardware and software develop best together, as they do in the lab of Christine P. Hendon, associate professor of electrical engineering. She and her students refine near-infrared spectroscopy (NIRS) and optical coherence tomography (OCT), both of which emit light into the body and measure what reflects. “By using fiber optics,” she says, “we can bring the light to wherever we need to within the body.”
One place they bring it to is the heart. Sometimes a small bit of tissue causes irregular beating, so doctors apply energy with a catheter to destroy the tissue. But there’s no good way to assess the depth of the lesion they create. Using ground-truth depth measures from dissection of animal tissue, donated human hearts, and live pigs, Hendon trains a deep-learning algorithm to predict depth from NIRS or OCT signals. “The beauty of neural networks is they can extract image features that are common across these cases,” Hendon says. Next, she hopes algorithms will not only detect lesion depth but create 2D maps of the tissue. In other work, she uses OCT to characterize the heart, detecting heart attack scarring or fat where there should be muscle. Compared with older machine learning techniques, accuracy has improved from around 80% to greater than 95%.
Hendon’s algorithms may help doctors evaluate and treat patients, and also shed light on the mechanisms of arrhythmia. She works closely with clinicians, collecting data, teaching them to use the new tools, and developing visualization methods to make the tools even more useful. “Seeing the images is very exciting to them,” she says.